| random_graph | Generate a random graph, with a given degree distribution and (optionally) vertex-vertex correlation. |
| random_rewire | Shuffle the graph in-place, following a variety of possible statistical models, chosen via the parameter model. |
| predecessor_tree | Return a graph from a list of predecessors given by the pred_map vertex property. |
| line_graph | Return the line graph of the given graph g. |
| graph_union | Return the union of graphs g1 and g2, composed of all edges and vertices of g1 and g2, without overlap. |
| triangulation | Generate a 2D or 3D triangulation graph from a given point set. |
| lattice | Generate a N-dimensional square lattice. |
| geometric_graph | Generate a geometric network form a set of N-dimensional points. |
| price_network | A generalized version of Price’s – or Barabási-Albert if undirected – preferential attachment network model. |
| complete_graph | Generate complete graph. |
| circular_graph | Generate a circular graph. |
Generate a random graph, with a given degree distribution and (optionally) vertex-vertex correlation.
The graph will be randomized via the random_rewire() function, and any remaining parameters will be passed to that function. Please read its documentation for all the options regarding the different statistical models which can be chosen.
| Parameters : | N : int
deg_sampler : function
directed : bool (optional, default: True)
parallel_edges : bool (optional, default: False)
self_loops : bool (optional, default: False)
block_membership : list or ndarray or function (optional, default: None)
block_type : string (optional, default: "int")
degree_block : bool (optional, default: False)
random : bool (optional, default: True)
verbose : bool (optional, default: False)
|
|---|---|
| Returns : | random_graph : Graph
blocks : PropertyMap
|
See also
Notes
The algorithm makes sure the degree sequence is graphical (i.e. realizable) and keeps re-sampling the degrees if is not. With a valid degree sequence, the edges are placed deterministically, and later the graph is shuffled with the random_rewire() function, with all remaining parameters passed to it.
The complexity is \(O(V + E)\) if parallel edges are allowed, and \(O(V + E \times\text{n-iter})\) if parallel edges are not allowed.
Note
If parallel_edges == False this algorithm only guarantees that the returned graph will be a random sample from the desired ensemble if n_iter is sufficiently large. The algorithm implements an efficient Markov chain based on edge swaps, with a mixing time which depends on the degree distribution and correlations desired. If degree correlations are provided, the mixing time tends to be larger.
References
| [metropolis-equations-1953] | Metropolis, N.; Rosenbluth, A.W.; Rosenbluth, M.N.; Teller, A.H.; Teller, E. “Equations of State Calculations by Fast Computing Machines”. Journal of Chemical Physics 21 (6): 1087-1092 (1953). DOI: 10.1063/1.1699114 |
| [hastings-monte-carlo-1970] | Hastings, W.K. “Monte Carlo Sampling Methods Using Markov Chains and Their Applications”. Biometrika 57 (1): 97-109 (1970). DOI: 10.1093/biomet/57.1.97 |
| [holland-stochastic-1983] | Paul W. Holland, Kathryn Blackmond Laskey, and Samuel Leinhardt, “Stochastic blockmodels: First steps,” Social Networks 5, no. 2: 109-13 (1983) DOI: 10.1016/0378-8733(83)90021-7 |
| [karrer-stochastic-2011] | Brian Karrer and M. E. J. Newman, “Stochastic blockmodels and community structure in networks,” Physical Review E 83, no. 1: 016107 (2011) DOI: 10.1103/PhysRevE.83.016107 arXiv: 1008.3926 |
Examples
This is a degree sampler which uses rejection sampling to sample from the distribution \(P(k)\propto 1/k\), up to a maximum.
>>> def sample_k(max):
... accept = False
... while not accept:
... k = randint(1,max+1)
... accept = random() < 1.0/k
... return k
...
The following generates a random undirected graph with degree distribution \(P(k)\propto 1/k\) (with k_max=40) and an assortative degree correlation of the form:
>>> g = gt.random_graph(1000, lambda: sample_k(40), model="probabilistic",
... vertex_corr=lambda i, k: 1.0 / (1 + abs(i - k)), directed=False,
... n_iter=100)
>>> gt.scalar_assortativity(g, "out")
(0.6285094791115295, 0.010745128857935755)
The following samples an in,out-degree pair from the joint distribution:
with \(m_1 = 4\) and \(m_2 = 20\).
>>> def deg_sample():
... if random() > 0.5:
... return poisson(4), poisson(4)
... else:
... return poisson(20), poisson(20)
...
The following generates a random directed graph with this distribution, and plots the combined degree correlation.
>>> g = gt.random_graph(20000, deg_sample)
>>>
>>> hist = gt.combined_corr_hist(g, "in", "out")
>>>
>>> clf()
>>> imshow(hist[0].T, interpolation="nearest", origin="lower")
<...>
>>> colorbar()
<...>
>>> xlabel("in-degree")
<...>
>>> ylabel("out-degree")
<...>
>>> savefig("combined-deg-hist.pdf")
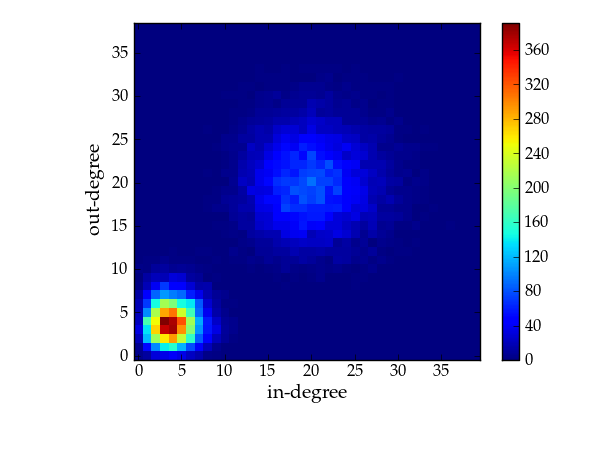
Combined degree histogram.
A correlated directed graph can be build as follows. Consider the following degree correlation:
i.e., the in->out correlation is “disassortative”, the out->in correlation is “assortative”, and everything else is uncorrelated. We will use a flat degree distribution in the range [1,20).
>>> p = scipy.stats.poisson
>>> g = gt.random_graph(20000, lambda: (sample_k(19), sample_k(19)),
... model="probabilistic",
... vertex_corr=lambda a,b: (p.pmf(a[0], b[1]) *
... p.pmf(a[1], 20 - b[0])),
... n_iter=100)
Lets plot the average degree correlations to check.
>>> clf()
>>> axes([0.1,0.15,0.63,0.8])
<...>
>>> corr = gt.avg_neighbour_corr(g, "in", "in")
>>> errorbar(corr[2][:-1], corr[0], yerr=corr[1], fmt="o-",
... label=r"$\left<\text{in}\right>$ vs in")
<...>
>>> corr = gt.avg_neighbour_corr(g, "in", "out")
>>> errorbar(corr[2][:-1], corr[0], yerr=corr[1], fmt="o-",
... label=r"$\left<\text{out}\right>$ vs in")
<...>
>>> corr = gt.avg_neighbour_corr(g, "out", "in")
>>> errorbar(corr[2][:-1], corr[0], yerr=corr[1], fmt="o-",
... label=r"$\left<\text{in}\right>$ vs out")
<...>
>>> corr = gt.avg_neighbour_corr(g, "out", "out")
>>> errorbar(corr[2][:-1], corr[0], yerr=corr[1], fmt="o-",
... label=r"$\left<\text{out}\right>$ vs out")
<...>
>>> legend(bbox_to_anchor=(1.01, 0.5), loc="center left", borderaxespad=0.)
<...>
>>> xlabel("Source degree")
<...>
>>> ylabel("Average target degree")
<...>
>>> savefig("deg-corr-dir.pdf")
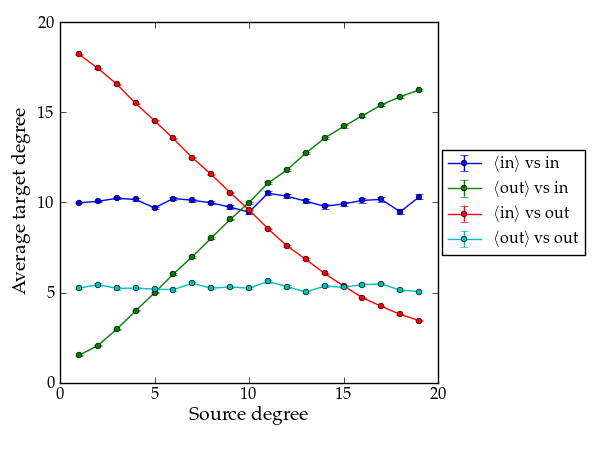
Average nearest neighbour correlations.
Stochastic blockmodels
The following example shows how a stochastic blockmodel [holland-stochastic-1983] [karrer-stochastic-2011] can be generated. We will consider a system of 10 blocks, which form communities. The connection probability will be given by
>>> def corr(a, b):
... if a == b:
... return 0.999
... else:
... return 0.001
The blockmodel can be generated as follows.
>>> g, bm = gt.random_graph(5000, lambda: poisson(10), directed=False,
... model="blockmodel-traditional",
... block_membership=lambda: randint(10),
... vertex_corr=corr)
>>> gt.graph_draw(g, vertex_fill_color=bm, edge_color="black", output="blockmodel.pdf")
<...>
Simple blockmodel with 10 blocks.
Shuffle the graph in-place, following a variety of possible statistical models, chosen via the parameter model.
| Parameters : | g : Graph
model : string (optional, default: "uncorrelated")
n_iter : int (optional, default: 1)
edge_sweep : bool (optional, default: True)
parallel : bool (optional, default: False)
self_loops : bool (optional, default: False)
vertex_corr : function (optional, default: None)
block_membership : PropertyMap (optional, default: None)
alias : bool (optional, default: True)
cache_probs : bool (optional, default: True)
persist : bool (optional, default: False)
verbose : bool (optional, default: False)
|
|---|---|
| Returns : | rejection_count : int
|
See also
Notes
This algorithm iterates through all the edges in the network and tries to swap its target or source with the target or source of another edge. The selected canditate swaps are chosen according to the model parameter.
Note
If parallel_edges = False, parallel edges are not placed during rewiring. In this case, the returned graph will be a uncorrelated sample from the desired ensemble only if n_iter is sufficiently large. The algorithm implements an efficient Markov chain based on edge swaps, with a mixing time which depends on the degree distribution and correlations desired. If degree probabilistic correlations are provided, the mixing time tends to be larger.
If model is either “probabilistic” or “blockmodel”, the Markov chain still needs to be mixed, even if parallel edges and self-loops are allowed. In this case the Markov chain is implemented using the Metropolis-Hastings [metropolis-equations-1953] [hastings-monte-carlo-1970] acceptance/rejection algorithm. It will eventually converge to the desired probabilities for sufficiently large values of n_iter.
Each edge is tentatively swapped once per iteration, so the overall complexity is \(O(V + E \times \text{n-iter})\). If edge_sweep == False, the complexity becomes \(O(V + E + \text{n-iter})\).
References
| [metropolis-equations-1953] | Metropolis, N.; Rosenbluth, A.W.; Rosenbluth, M.N.; Teller, A.H.; Teller, E. “Equations of State Calculations by Fast Computing Machines”. Journal of Chemical Physics 21 (6): 1087-1092 (1953). DOI: 10.1063/1.1699114 |
| [hastings-monte-carlo-1970] | Hastings, W.K. “Monte Carlo Sampling Methods Using Markov Chains and Their Applications”. Biometrika 57 (1): 97-109 (1970). DOI: 10.1093/biomet/57.1.97 |
| [holland-stochastic-1983] | Paul W. Holland, Kathryn Blackmond Laskey, and Samuel Leinhardt, “Stochastic blockmodels: First steps,” Social Networks 5, no. 2: 109-13 (1983) DOI: 10.1016/0378-8733(83)90021-7 |
| [karrer-stochastic-2011] | Brian Karrer and M. E. J. Newman, “Stochastic blockmodels and community structure in networks,” Physical Review E 83, no. 1: 016107 (2011) DOI: 10.1103/PhysRevE.83.016107 arXiv: 1008.3926 |
Examples
Some small graphs for visualization.
>>> g, pos = gt.triangulation(random((1000,2)))
>>> pos = gt.arf_layout(g)
>>> gt.graph_draw(g, pos=pos, output="rewire_orig.pdf", output_size=(300, 300))
<...>
>>> gt.random_rewire(g, "correlated")
189
>>> pos = gt.arf_layout(g)
>>> gt.graph_draw(g, pos=pos, output="rewire_corr.pdf", output_size=(300, 300))
<...>
>>> gt.random_rewire(g)
197
>>> pos = gt.arf_layout(g)
>>> gt.graph_draw(g, pos=pos, output="rewire_uncorr.pdf", output_size=(300, 300))
<...>
>>> gt.random_rewire(g, "erdos")
26
>>> pos = gt.arf_layout(g)
>>> gt.graph_draw(g, pos=pos, output="rewire_erdos.pdf", output_size=(300, 300))
<...>
Some ridiculograms :




From left to right: Original graph; Shuffled graph, with degree correlations; Shuffled graph, without degree correlations; Shuffled graph, with random degrees.
We can try with larger graphs to get better statistics, as follows.
>>> figure()
<...>
>>> g = gt.random_graph(30000, lambda: sample_k(20), model="probabilistic",
... vertex_corr=lambda i, j: exp(abs(i-j)), directed=False,
... n_iter=100)
>>> corr = gt.avg_neighbour_corr(g, "out", "out")
>>> errorbar(corr[2][:-1], corr[0], yerr=corr[1], fmt="o-", label="Original")
<...>
>>> gt.random_rewire(g, "correlated")
206
>>> corr = gt.avg_neighbour_corr(g, "out", "out")
>>> errorbar(corr[2][:-1], corr[0], yerr=corr[1], fmt="*", label="Correlated")
<...>
>>> gt.random_rewire(g)
109
>>> corr = gt.avg_neighbour_corr(g, "out", "out")
>>> errorbar(corr[2][:-1], corr[0], yerr=corr[1], fmt="o-", label="Uncorrelated")
<...>
>>> gt.random_rewire(g, "erdos")
13
>>> corr = gt.avg_neighbour_corr(g, "out", "out")
>>> errorbar(corr[2][:-1], corr[0], yerr=corr[1], fmt="o-", label=r"Erd\H{o}s")
<...>
>>> xlabel("$k$")
<...>
>>> ylabel(r"$\left<k_{nn}\right>$")
<...>
>>> legend(loc="best")
<...>
>>> savefig("shuffled-stats.pdf")
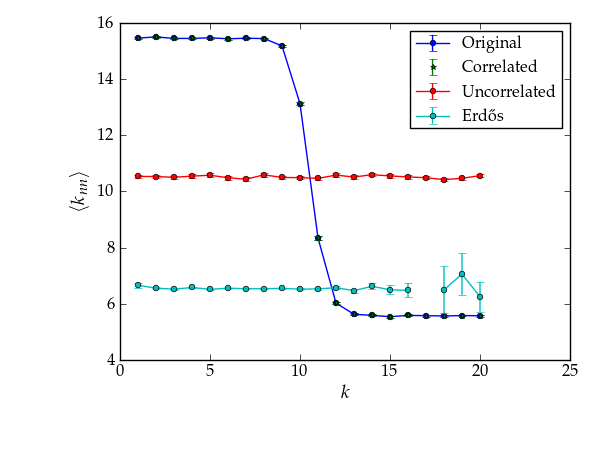
Average degree correlations for the different shuffled and non-shuffled graphs. The shuffled graph with correlations displays exactly the same correlation as the original graph.
Now let’s do it for a directed graph. See random_graph() for more details.
>>> p = scipy.stats.poisson
>>> g = gt.random_graph(20000, lambda: (sample_k(19), sample_k(19)),
... model="probabilistic",
... vertex_corr=lambda a, b: (p.pmf(a[0], b[1]) * p.pmf(a[1], 20 - b[0])),
... n_iter=100)
>>> figure()
<...>
>>> axes([0.1,0.15,0.6,0.8])
<...>
>>> corr = gt.avg_neighbour_corr(g, "in", "out")
>>> errorbar(corr[2][:-1], corr[0], yerr=corr[1], fmt="o-",
... label=r"$\left<\text{o}\right>$ vs i")
<...>
>>> corr = gt.avg_neighbour_corr(g, "out", "in")
>>> errorbar(corr[2][:-1], corr[0], yerr=corr[1], fmt="o-",
... label=r"$\left<\text{i}\right>$ vs o")
<...>
>>> gt.random_rewire(g, "correlated")
4323
>>> corr = gt.avg_neighbour_corr(g, "in", "out")
>>> errorbar(corr[2][:-1], corr[0], yerr=corr[1], fmt="o-",
... label=r"$\left<\text{o}\right>$ vs i, corr.")
<...>
>>> corr = gt.avg_neighbour_corr(g, "out", "in")
>>> errorbar(corr[2][:-1], corr[0], yerr=corr[1], fmt="o-",
... label=r"$\left<\text{i}\right>$ vs o, corr.")
<...>
>>> gt.random_rewire(g, "uncorrelated")
153
>>> corr = gt.avg_neighbour_corr(g, "in", "out")
>>> errorbar(corr[2][:-1], corr[0], yerr=corr[1], fmt="o-",
... label=r"$\left<\text{o}\right>$ vs i, uncorr.")
<...>
>>> corr = gt.avg_neighbour_corr(g, "out", "in")
>>> errorbar(corr[2][:-1], corr[0], yerr=corr[1], fmt="o-",
... label=r"$\left<\text{i}\right>$ vs o, uncorr.")
<...>
>>> legend(bbox_to_anchor=(1.01, 0.5), loc="center left", borderaxespad=0.)
<...>
>>> xlabel("Source degree")
<...>
>>> ylabel("Average target degree")
<...>
>>> savefig("shuffled-deg-corr-dir.pdf")
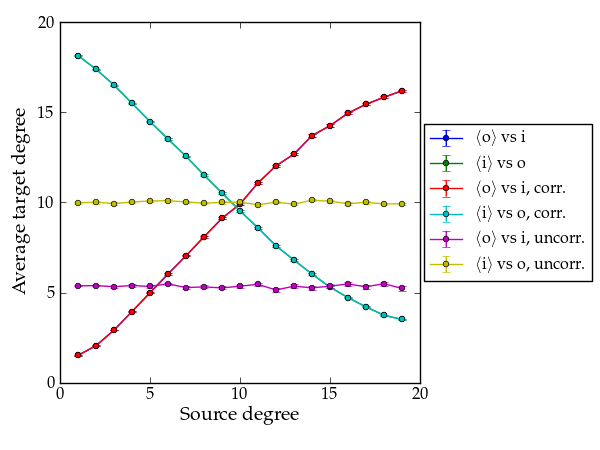
Average degree correlations for the different shuffled and non-shuffled directed graphs. The shuffled graph with correlations displays exactly the same correlation as the original graph.
Return a graph from a list of predecessors given by the pred_map vertex property.
Return the line graph of the given graph g.
Notes
Given an undirected graph G, its line graph L(G) is a graph such that
- each vertex of L(G) represents an edge of G; and
- two vertices of L(G) are adjacent if and only if their corresponding edges share a common endpoint (“are adjacent”) in G.
For a directed graph, the second criterion becomes:
- Two vertices representing directed edges from u to v and from w to x in G are connected by an edge from uv to wx in the line digraph when v = w.
References
| [line-wiki] | http://en.wikipedia.org/wiki/Line_graph |
Examples
>>> g = gt.collection.data["lesmis"]
>>> lg, vmap = gt.line_graph(g)
>>> gt.graph_draw(g, pos=g.vp["pos"], output="lesmis.pdf")
<...>
>>> pos = gt.graph_draw(lg, output="lesmis-lg.pdf")
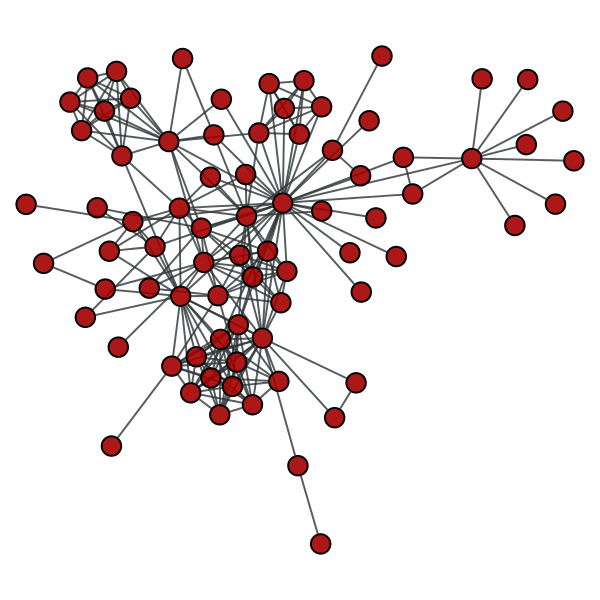
Coappearances of characters in Victor Hugo’s novel “Les Miserables”.
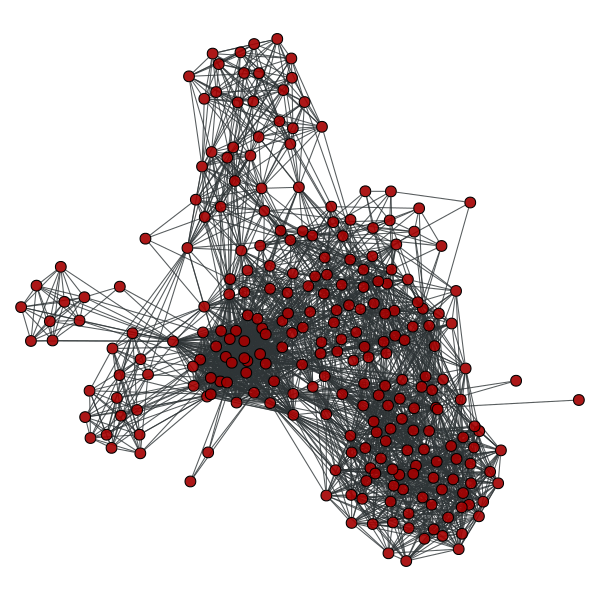
Line graph of the coappearance network on the left.
Return the union of graphs g1 and g2, composed of all edges and vertices of g1 and g2, without overlap.
| Parameters : | g1 : Graph
g2 : Graph
intersection : PropertyMap (optional, default: None)
props : list of tuples of PropertyMap (optional, default: [])
include : bool (optional, default: False)
|
|---|---|
| Returns : | ug : Graph
props : list of PropertyMap objects
|
Examples
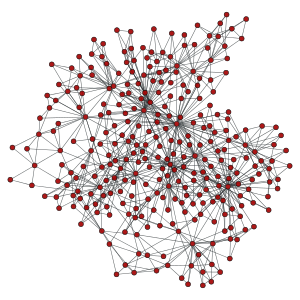
>>> g = gt.triangulation(random((300,2)))[0]
>>> ug = gt.graph_union(g, g)
>>> uug = gt.graph_union(g, ug)
>>> pos = gt.sfdp_layout(g)
>>> gt.graph_draw(g, pos=pos, output_size=(300,300), output="graph_original.pdf")
<...>
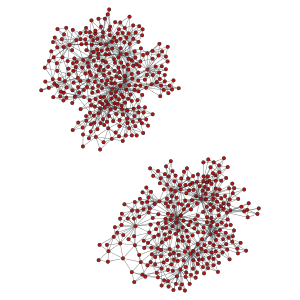
>>> pos = gt.sfdp_layout(ug)
>>> gt.graph_draw(ug, pos=pos, output_size=(300,300), output="graph_union.pdf")
<...>
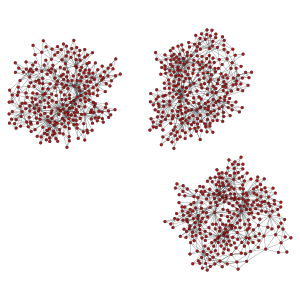
>>> pos = gt.sfdp_layout(uug)
>>> gt.graph_draw(uug, pos=pos, output_size=(300,300), output="graph_union2.pdf")
<...>



Generate a 2D or 3D triangulation graph from a given point set.
| Parameters : | points : ndarray
type : string (optional, default: 'simple')
periodic : bool (optional, default: False)
|
|---|---|
| Returns : | triangulation_graph : Graph
pos : PropertyMap
|
See also
Notes
A triangulation [cgal-triang] is a division of the convex hull of a point set into triangles, using only that set as triangle vertices.
In simple triangulations (type=”simple”), the insertion of a point is done by locating a face that contains the point, and splitting this face into three new faces (the order of insertion is therefore important). If the point falls outside the convex hull, the triangulation is restored by flips. Apart from the location, insertion takes a time O(1). This bound is only an amortized bound for points located outside the convex hull.
Delaunay triangulations (type=”delaunay”) have the specific empty sphere property, that is, the circumscribing sphere of each cell of such a triangulation does not contain any other vertex of the triangulation in its interior. These triangulations are uniquely defined except in degenerate cases where five points are co-spherical. Note however that the CGAL implementation computes a unique triangulation even in these cases.
References
| [cgal-triang] | (1, 2) http://www.cgal.org/Manual/last/doc_html/cgal_manual/Triangulation_3/Chapter_main.html |
Examples
>>> points = random((500, 2)) * 4
>>> g, pos = gt.triangulation(points)
>>> weight = g.new_edge_property("double") # Edge weights corresponding to
... # Euclidean distances
>>> for e in g.edges():
... weight[e] = sqrt(sum((array(pos[e.source()]) -
... array(pos[e.target()]))**2))
>>> b = gt.betweenness(g, weight=weight)
>>> b[1].a *= 100
>>> gt.graph_draw(g, pos=pos, output_size=(300,300), vertex_fill_color=b[0],
... edge_pen_width=b[1], output="triang.pdf")
<...>
>>> g, pos = gt.triangulation(points, type="delaunay")
>>> weight = g.new_edge_property("double")
>>> for e in g.edges():
... weight[e] = sqrt(sum((array(pos[e.source()]) -
... array(pos[e.target()]))**2))
>>> b = gt.betweenness(g, weight=weight)
>>> b[1].a *= 120
>>> gt.graph_draw(g, pos=pos, output_size=(300,300), vertex_fill_color=b[0],
... edge_pen_width=b[1], output="triang-delaunay.pdf")
<...>
2D triangulation of random points:
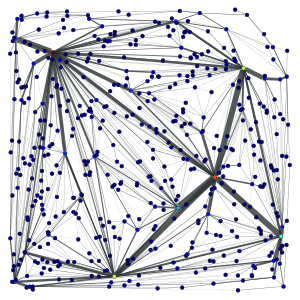
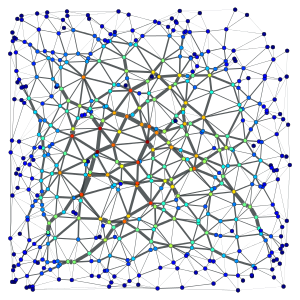


Left: Simple triangulation. Right: Delaunay triangulation. The vertex colors and the edge thickness correspond to the weighted betweenness centrality.
Generate a N-dimensional square lattice.
| Parameters : | shape : list or ndarray
periodic : bool (optional, default: False)
|
|---|---|
| Returns : | lattice_graph : Graph
|
See also
References
| [lattice] | http://en.wikipedia.org/wiki/Square_lattice |
Examples
>>> g = gt.lattice([10,10])
>>> pos = gt.sfdp_layout(g, cooling_step=0.95, epsilon=1e-2)
>>> gt.graph_draw(g, pos=pos, output_size=(300,300), output="lattice.pdf")
<...>
>>> g = gt.lattice([10,20], periodic=True)
>>> pos = gt.sfdp_layout(g, cooling_step=0.95, epsilon=1e-2)
>>> gt.graph_draw(g, pos=pos, output_size=(300,300), output="lattice_periodic.pdf")
<...>
>>> g = gt.lattice([10,10,10])
>>> pos = gt.sfdp_layout(g, cooling_step=0.95, epsilon=1e-2)
>>> gt.graph_draw(g, pos=pos, output_size=(300,300), output="lattice_3d.pdf")
<...>
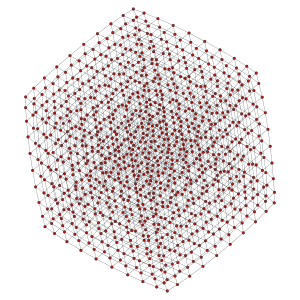
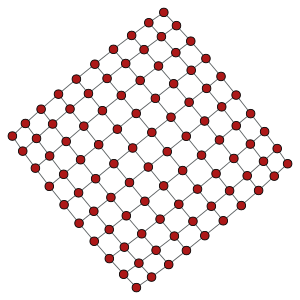
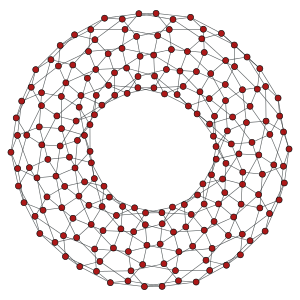

Left: 10x10 2D lattice. Middle: 10x20 2D periodic lattice (torus). Right: 10x10x10 3D lattice.
Generate complete graph.
| Parameters : | N : int
self_loops : bool (optional, default: False)
directed : bool (optional, default: False)
|
|---|---|
| Returns : | complete_graph : Graph
|
References
| [complete] | http://en.wikipedia.org/wiki/Complete_graph |
Examples
>>> g = gt.complete_graph(30)
>>> pos = gt.sfdp_layout(g, cooling_step=0.95, epsilon=1e-2)
>>> gt.graph_draw(g, pos=pos, output_size=(300,300), output="complete.pdf")
<...>
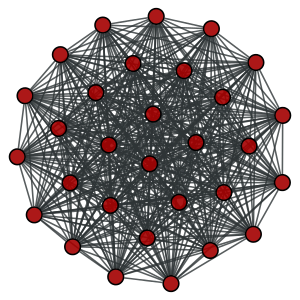
A complete graph with \(N=30\) vertices.
Generate a circular graph.
| Parameters : | N : int
k : int (optional, default: True)
self_loops : bool (optional, default: False)
directed : bool (optional, default: False)
|
|---|---|
| Returns : | circular_graph : Graph
|
Examples
>>> g = gt.circular_graph(30, 2)
>>> pos = gt.sfdp_layout(g, cooling_step=0.95)
>>> gt.graph_draw(g, pos=pos, output_size=(300,300), output="circular.pdf")
<...>
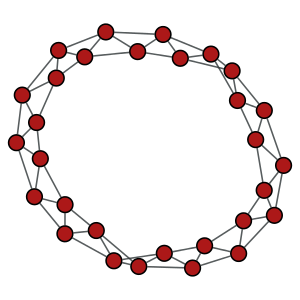
A circular graph with \(N=30\) vertices, and \(k=2\).
Generate a geometric network form a set of N-dimensional points.
| Parameters : | points : list or ndarray
radius : float
ranges : list or ndarray (optional, default: None)
|
|---|---|
| Returns : | geometric_graph : Graph
pos : PropertyMap
|
See also
Notes
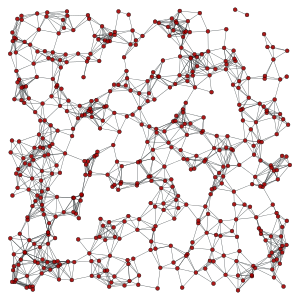
A geometric graph [geometric-graph] is generated by connecting points embedded in a N-dimensional euclidean space which are at a distance equal to or smaller than a given radius.
References
| [geometric-graph] | (1, 2) Jesper Dall and Michael Christensen, “Random geometric graphs”, Phys. Rev. E 66, 016121 (2002), DOI: 10.1103/PhysRevE.66.016121 |
Examples
>>> points = random((500, 2)) * 4
>>> g, pos = gt.geometric_graph(points, 0.3)
>>> gt.graph_draw(g, pos=pos, output_size=(300,300), output="geometric.pdf")
<...>
>>> g, pos = gt.geometric_graph(points, 0.3, [(0,4), (0,4)])
>>> pos = gt.graph_draw(g, output_size=(300,300), output="geometric_periodic.pdf")
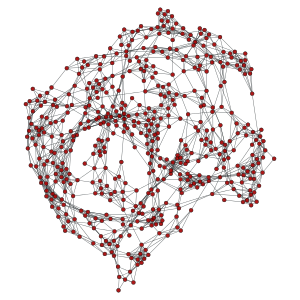

A generalized version of Price’s – or Barabási-Albert if undirected – preferential attachment network model.
| Parameters : | N : int
m : int (optional, default: 1)
c : float (optional, default: 1 if directed == True else 0)
gamma : float (optional, default: 1)
directed : bool (optional, default: True)
seed_graph : Graph (optional, default: None)
|
|---|---|
| Returns : | price_graph : Graph
|
See also
Notes
The (generalized) [price] network is either a directed or undirected graph (the latter is called a Barabási-Albert network), generated dynamically by at each step adding a new vertex, and connecting it to \(m\) other vertices, chosen with probability \(\pi\) defined as:
where \(k\) is the in-degree of the vertex (or simply the degree in the undirected case). If \(\gamma=1\), the tail of resulting in-degree distribution of the directed case is given by
or for the undirected case
However, if \(\gamma \ne 1\), the in-degree distribution is not scale-free (see [dorogovtsev-evolution] for details).
Note that if seed_graph is not given, the algorithm will always start with one node if \(c > 0\), or with two nodes with a link between them otherwise. If \(m > 1\), the degree of the newly added vertices will be vary dynamically as \(m'(t) = \min(m, N(t))\), where \(N(t)\) is the number of vertices added so far. If this behaviour is undesired, a proper seed graph with \(N \ge m\) vertices must be provided.
This algorithm runs in \(O(N\log N)\) time.
References
| [yule] | Yule, G. U. “A Mathematical Theory of Evolution, based on the Conclusions of Dr. J. C. Willis, F.R.S.”. Philosophical Transactions of the Royal Society of London, Ser. B 213: 21-87, 1925, DOI: 10.1098/rstb.1925.0002 |
| [price] | (1, 2) Derek De Solla Price, “A general theory of bibliometric and other cumulative advantage processes”, Journal of the American Society for Information Science, Volume 27, Issue 5, pages 292-306, September 1976, DOI: 10.1002/asi.4630270505 |
| [barabasi-albert] | Barabási, A.-L., and Albert, R., “Emergence of scaling in random networks”, Science, 286, 509, 1999, DOI: 10.1126/science.286.5439.509 |
| [dorogovtsev-evolution] | (1, 2) S. N. Dorogovtsev and J. F. F. Mendes, “Evolution of networks”, Advances in Physics, 2002, Vol. 51, No. 4, 1079-1187, DOI: 10.1080/00018730110112519 |
Examples
>>> g = gt.price_network(100000)
>>> gt.graph_draw(g, pos=gt.sfdp_layout(g, epsilon=1e-2, cooling_step=0.95),
... vertex_fill_color=g.vertex_index, vertex_size=2,
... edge_pen_width=1, output="price-network.png")
<...>
>>> g = gt.price_network(100000, c=0.1)
>>> gt.graph_draw(g, pos=gt.sfdp_layout(g, epsilon=1e-2, cooling_step=0.95),
... vertex_fill_color=g.vertex_index, vertex_size=2,
... edge_pen_width=1, output="price-network-broader.png")
<...>
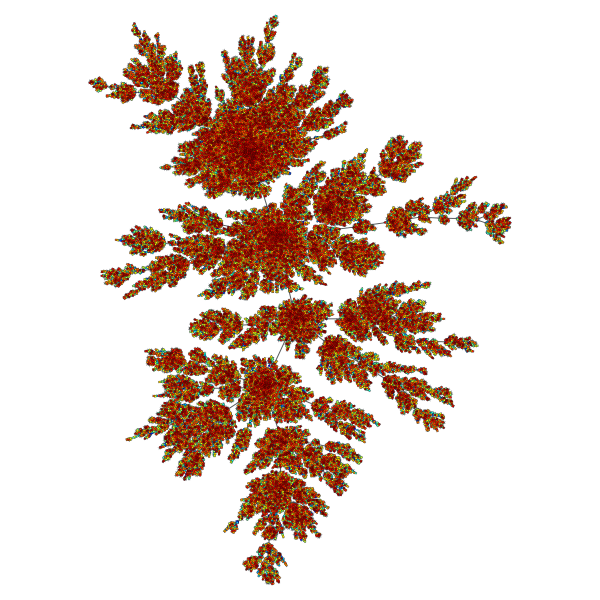

Price network with \(N=10^5\) nodes and \(c=1\). The colors represent the order in which vertices were added.
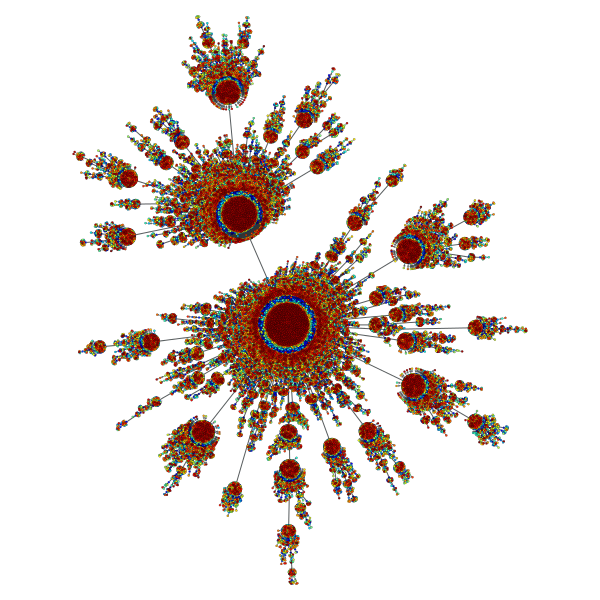

Price network with \(N=10^5\) nodes and \(c=0.1\). The colors represent the order in which vertices were added.